scMultiSim
Author
Assistant Professor, Georgia Institute of Technology
代表工作(来自课题组工作网站):
- Single cell multi-modal, multi-batch, multi-condition data integration and analysis. (Example projects: scDART, scMoMaT,scDisInFact)
- Temporal analysis of single cells: how cells change over pseudotime or real time through cell divisions. (Example projects: CellPath, LinRace)
- Spatial-omics and Spatial-temporal dynamics of cells (Example projects: scHybridNMF, CLARIFY, TemSOMap, SpaDecoder)
- Simulation of single cell omics (including temporal and spatial) data to evaluation computational methods. (Example projects: TedSim, scMultiSim)
- Inference of gene regulatory networks, cell-cell interactions, and cross-modality relationships in multi-omics data. (Example projects: CespGRN, CLARIFY)
Simulation of single cell omics
To evaluate the performance of proposed computational methods
By generating data that models biological mechanisms and provides ground truth for benchmarking
一句话概括:实验数据缺失或不足的情况下,模拟数据→评估计算方法
| Tools | id | GRNs | velocity | CCI | reference-based | multiomic |
|---|---|---|---|---|---|---|
| SymSim | ✔️ | |||||
| SERGIO | ✔️ | ✔️ | ✔️❌ | |||
| BEELINE | ✔️ | |||||
| dyngen | ✔️ | ✔️ | ✔️❌ | |||
| VeloSim | ✔️ | |||||
| mistyR | ✔️ | |||||
| scDesign3 | ✔️ | ✔️ |
CCI: cell–cell interaction
Methods
模型介绍
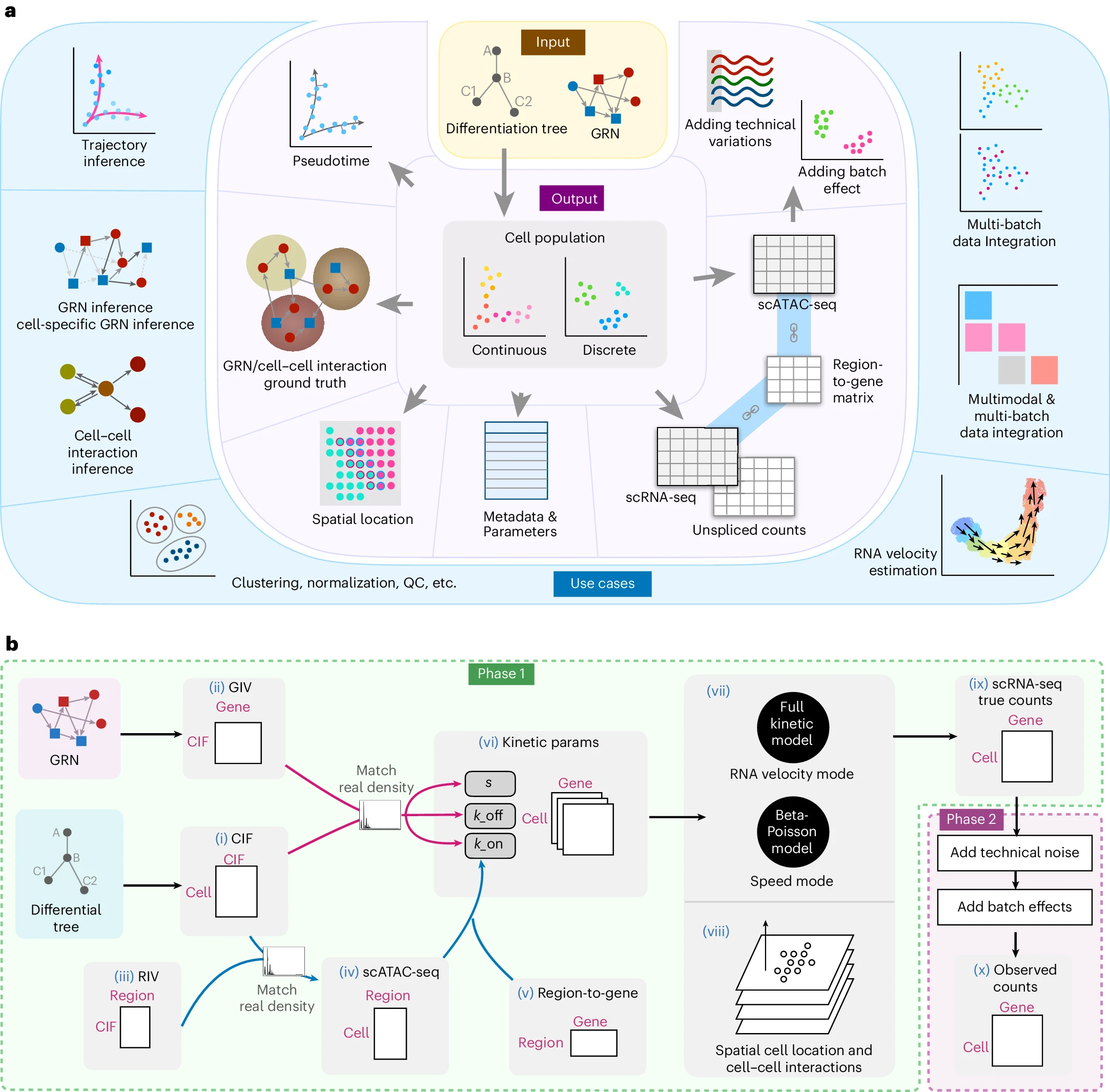
| Input | Output |
|---|---|
| cell identity,RNA velocity, GRNs, CCI, relationships between chromatin accessibility and transcriptome data |
①能够解决的问题:
测评 clustering, trajectory inference, RNA velocity estimation, multimodal and multi-batch data integration, GRN inference and CCI inference
mosaic integration,特指 multimodal and multi-batch data integration
②主要创新点:
using multi-omics data without using only scRNA-seq data
allows users to adjust the effect of each biological factor on the output data→investigate how the methods’ performance is affected by each factor
原理
Single cell multi-modal, multi-batch, multi-condition data integration and analysis. (Example projects: scDART, scMoMaT,scDisInFact)
Temporal analysis of single cells: how cells change over pseudotime or real time through cell divisions. (Example projects: CellPath, LinRace)
Spatial-omics and Spatial-temporal dynamics of cells (Example projects: scHybridNMF, CLARIFY, TemSOMap, SpaDecoder)
Simulation of single cell omics (including temporal and spatial) data to evaluation computational methods. (Example projects: TedSim, scMultiSim)
Inference of gene regulatory networks, cell-cell interactions, and cross-modality relationships in multi-omics data. (Example projects: CespGRN, CLARIFY)

